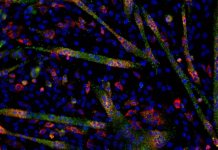
Understand aging with genomics, proteomics, and all things -omics!
“Omics” approaches are a set of analytical tools to explore the functions of our bodies. The best known ones are genomics, transcriptomics, proteomics and metabolomics, but newer branches have recently emerged, such as lipidomics, glycomics or nutriomics.
Genomics correspond to the study of the genome, i.e. everything that concerns DNA modifications, mutations, expression… Transcriptomics take a look at the next stage, everything in the order of the transcriptome, messenger RNAs coding for our proteins, non-coding RNAs, transcription regulation… For proteomics, it is still the next step, namely the study of proteins, their translation, folding and modification. Finally, metabolomics is the analysis of chemical factors regulating inter- and intracellular interactions, also called metabolites.
New types of analysis include the study of lipids (lipidomics), carbohydrates (glycomics) and nutrition and its impact on our bodies (nutriomics). These are fields as vast as the first and which constitute the future of research in “omics” approaches.
Fight aging with multifactorial “-omics” studies
Who hasn’t heard of telomeres when we’re interested in aging? Epigenetic regulation phenomena? Sirtuins? If at least one of these themes speaks to you, it is because you have already set foot in the “omics” approaches, perhaps without knowing it!
Genomics, unlike genetics which studies genes one by one in a given individual or small population and their role in offspring, focuses on the analysis of the structure, function and editing of the genome as a whole. With this technique, it is possible to account for the overall function of a cell and its potential dysfunctions. Genomics also includes DNA sequencing, a technique increasingly used in disease diagnostics that, combined with bioinformatics, allows projections of the evolution of a cell or tissue. Thus, thanks to this tool, it is possible to determine the age of a cell but also its life expectancy, by measuring its telomeres for instance. This “omics” approach has also given rise to systems biology, thanks to which it is now possible to understand and model the function of complex organs.
Transcriptomics is a slightly more complicated approach, because it is an analysis that looks at all RNAs, coding or not. By coding we mean that an RNA will ultimately give a protein. The relatively recent discovery of non-coding RNAs has revolutionized this type of analysis, including previously unknown or misunderstood regulatory processes. As in genomics, it is possible to sequence the entire transcriptome, i.e. all the RNAs of a cell, including messenger RNAs (mRNAs that will give proteins), ribosomal RNAs (specific to ribosomes), Interfering RNAs (which interact with mRNAs leading to their degradation) and other non-coding RNAs (such as microRNAs, piwi-RNAs or nuclear RNAs, whose roles are still under study). All these data allow us to measure the expression of a gene in different tissues or conditions, thus giving us an overview of gene regulation, functions of a specific gene or changes in expression under pathological conditions.


Proteomics is more targeted and even less used than the two previous “omics” approaches. It is part of the study of the proteome, all the proteins of a cell. In these analyses, we can look at the changes that a protein undergoes during its synthesis: its translation, post-translational changes (such as acetylation, methylation…), its folding (the 3D structure of a protein being central to its function), its coupling with other proteins (formation of dimers, trimers or polymers ensuring adequate action of the protein in question) or its degradation. This discipline is mainly used to identify potential therapeutic targets or biomarkers of pathologies, but its applications are becoming more and more diversified, particularly on protein-DNA, protein-RNA and protein-protein interactions, thus joining genomics, transcriptomics and systems biology.
Metabolomics is based on the concept that each process taking place in a cell (the set of processes being called metabolism) leaves a chemical trace, before, during or after said process, in the form of a metabolite. By mapping the metabolites available in a cell, it is possible to account for its function and metabolism. This discipline is booming and requires important research, since the Human Metabolome Database lists about 3000 human metabolites at present, against nearly 50 000 in plants!
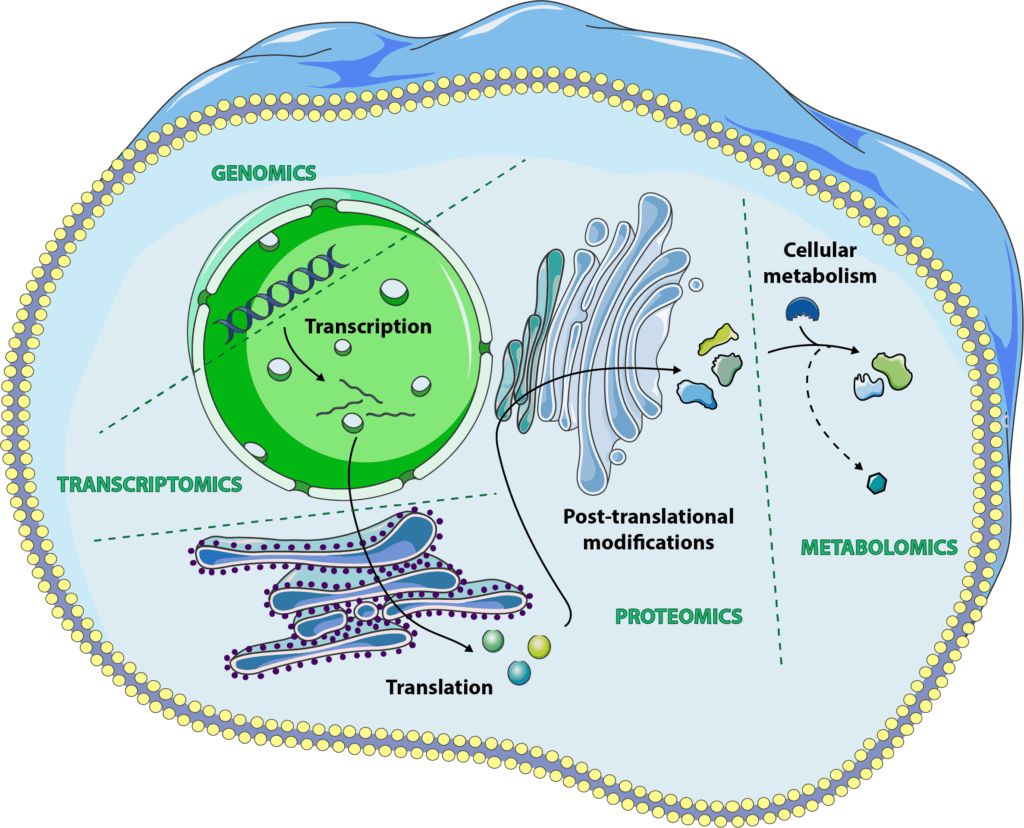
Combining all these “omics” approaches, it is now possible to have an assessment of one’s biological age and to study the processes leading to physiological aging and its associated diseases. In this series of articles, we will see in more detail how the different “omics” work, what analytical techniques they use and their various applications in the fight against aging.
See all our articles on the “-omics” approaches
The “-omics” : a tool to better understand our aging process
What is behind the term “-omics? When we talk of genomics, transcriptomics or proteomics, what are we looking at? Here is a guide to explain it all!
Part 1: Understanding genomics for anti-aging research

How can we not mention genomics and how useful they are. It is the oldest “-omics” approach and still the most studied one even today. It gave birth to whole new concepts, such as epigenetics or systems biology, and opened up the scientific community to new horizons that we had never even dreamed ot!
Part 2: Transcriptomics, a constantly evolving science.

The discovery of non coding RNA led to a Nobel Prize. Enough to say it’s an important field of research! Transcriptomics is allowing the discovery of new tools, new mechanisms and led us to better understand the regulation of transcription.
Part 3: Proteomics, a mish-mash of research fields
 Above all, proteomics are a multidisciplinary approach, taking into account the interactions between fields, namely genomics and transcriptomics. It also refers to different concepts, from immunology to nutrition or cell function.
Above all, proteomics are a multidisciplinary approach, taking into account the interactions between fields, namely genomics and transcriptomics. It also refers to different concepts, from immunology to nutrition or cell function.
Part 4: Metabolomics, the last-born of the “omics”
 Last but not least! Metabolomics is a field that helps us understand very complicated regulation networks and the daily discovery of new cell-cell communication players.
Last but not least! Metabolomics is a field that helps us understand very complicated regulation networks and the daily discovery of new cell-cell communication players.
Dr. Marion Tible

Author/Reviewer
Auteure/Relectrice
Marion Tible has a PhD in cellular biology and physiopathology. Formerly a researcher in thematics varying from cardiology to neurodegenerative diseases, she is now part of Long Long Life team and is involved in scientific writing and anti-aging research.
More about the Long Long Life team
Marion Tible est docteur en biologie cellulaire et physiopathologie. Ancienne chercheuse dans des thématiques oscillant de la cardiologie aux maladies neurodégénératives, elle est aujourd’hui impliquée au sein de Long Long Life pour la rédaction scientifique et la recherche contre le vieillissement.
En savoir plus sur l’équipe de Long Long Life
Dr Guilhem Velvé Casquillas

Author/Reviewer
Auteur/Relecteur
Physics PhD, CEO NBIC Valley, CEO Long Long Life, CEO Elvesys Microfluidic Innovation Center
More about the Long Long Life team
Docteur en physique, CEO NBIC Valley, CEO Long Long Life, CEO Elvesys Microfluidic Innovation Center
En savoir plus sur l’équipe de Long Long Life


![[Video] Eurosymposium on Healthy Ageing, Brussels, 2018 Eurosymposium on Healthy Aging](https://www.longlonglife.org/wp-content/uploads/2019/07/P1310252-218x150.jpg)