Genomics is the study of the genome, that is to say, the entirety of our DNA. Knowing that if we unfolded all the DNA contained in our body, that could cover 1000 times the distance from the Earth to the Sun, we can wonder how it is possible to analyze such an amount of data!
Genomics: how to analyze a genome

In 1953, Watson & Crick, two researchers basing their work on Wilkins & Franklin crystallographic data have succeeded in solving mystery of DNA structure, opening up a whole new field of study, especially concerning DNA replication and transcription processes. Sequencing was still far away but the idea took form among the researchers and in 1965, the first transcriptome was sequenced. It was that of a yeast, Saccharomyces cerevisiae[1]. However, though RNA is good, DNA is better! Technological advances have made it possible to develop different sequencing techniques, which won Gilbert & Sanger a Nobel Prize in chemistry in 1970. Finally, in 1977, the first genome to be fully sequenced belonged to a bacteriophage virus[2].
Hpw does one sequence a genome?
Sequencing by the Sanger method[3] – by enzymatic synthesis
It is the most widely used technique today and many improvements have been made since the 1970s. The principle is relatively simple and based on the same mechanism as natural DNA replication: a primer (a small DNA sequence) will initiate the synthesis of a DNA strand complementary to the one we want to sequence, a DNA polymerase will elongate this strand, then a mixture of deoxyribonucleotides (dATP, dCTP, dGTP and dTTP) and di-deoxyribonucleotides (ddATP, ddCTP, ddGTP or ddTTP) will be introduced into the mixture. Deoxyribonucleotides will be normally integrated and participate in DNA strand synthesis (and give adenine, cytosine, guanine and thymine, the four bases of DNA). On the other hand, the di-desoxyribonucleotides (homologues of deoxyribonucleotides but not having the chemical group necessary for the action of the DNA polymerase), will integrate into the strand during synthesis and cause it to break. Thus, if ddCTP has been introduced into the mixture, the strand being synthesized will cut at each cytosine. For complete sequencing, simply repeat with the remaining three di-deoxyribonucleotides. The experimenter will end up with DNA fragments of different lengths for which they will know exactly on which di-deoxyribonucleotide they were broken. Each fragment is analyzed and reordered according to its molecular weight, allowing the exact DNA sequence to be found.
Gilbert sequencing[4] – chemical degradation
Unlike Sanger’s method, the Gilbert method is based on DNA degradation: the goal is to achieve selective cuts through chemical processes breaking DNA into fragments, which will be put back in order later. Once the DNA of interest is marked with a radioactive tracer, it is denatured (double stranded to single stranded) and subjected to chemical modifications, developed by Gilbert himself, specific to each base constituting the DNA. At the modification site, the DNA is then cut and each fragment is analyzed and returned in the order of its molecular weight (as for the Sanger method).
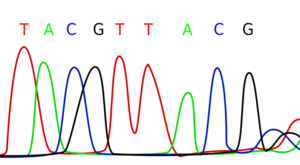
How to sequence an entire genome
Both of the above techniques have been greatly improved since their discovery, particularly in terms of sample preparation, separation and DNA fragment detection. These advances have made it possible to automate sequencing with increasingly efficient machines. However, the entire genome represents several billion bases and no machine today has the capacity to process so much information. It is necessary to prepare the DNA, by treating it with restriction enzymes or ultrasound, in order to obtain fragments that can be sequenced. The advent of bioinformatics and nanotechnologies has also revolutionized the field to create genomics as we know it today.
HTS, NGS, microarray, GWAS… What’s all this?
The latest developments in sequencing have given rise to many acronyms, not always easy to understand. HTS stands for High-Throughput Sequencing. It is often associated with NGS, “Next Generation Sequencing“, because the two approaches emerged in the early 2000s and overlap[5]. The aim of these techniques is to increase the number of decoded sequences per series of analyses, up to a few million DNA fragments, while lowering the cost of these tests. They have revolutionized genomics by making previously complex and expensive studies accessible in terms of time and money. They also make it possible, and this is a revolution in itself, to analyse a single copy of DNA, thus making it possible to analyse very small samples[5].
Pyrosequencing, which is part of HTS/NGS, is currently the most widely used: it is based on the Sanger method but uses pyrophosphate nucleotides instead of di-desoxyribonucleotides. When they are incorporated into the DNA strand during synthesis, instead of stopping the elongation, they will release a pyrophosphate that will be transformed and emits light. The result is a graph with peaks that show the incorporation of the marked nucleotides, and whose height reflects the number of nucleotides integrated into the DNA sequence. With bioinformatic tools, it is then possible to determine the DNA sequence. The major interest of this technique is the parallel analysis of several thousand fragments, allowing results to be obtained in a few hours[5].
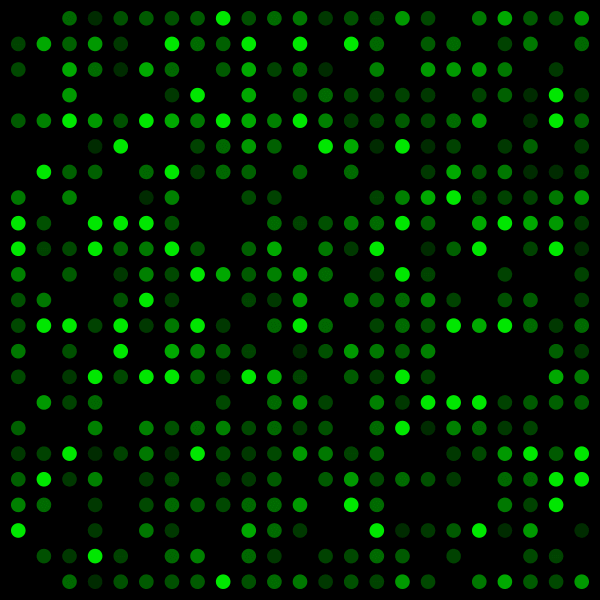

DNA chips, or “DNA-microarray“, use a completely different mechanism: known DNA sequences are attached to a plate (the chip) and the DNA to be tested is injected onto that surface. When it is complementary to a sequence on the chip, it hybridizes, emits detectable light and we obtain a hybridization spectrum that can give us several information: expression level of a given gene, mutations, sequencing, interactions with other molecules… [6]
The GWAS approach, “Genome-Wide Association Studies“, is a genetics technique crossed with genomics. As the name suggests, it studies genetic mutations, not at the level of a gene but by taking into account the entire genome. It is not a sequencing technique strictly speaking, but it allows mutations to be identified quickly and reliably[7]. It highlights mostly what are called SNPs, “single nucleotide polymorphism”, which correspond to mutations of a single base within a complete DNA sequence. It then links these SNPs to known diseases, thus linking the occurrence of a mutation to a pathological situation[7].
More recently, single-cell genomics have emerged, with the ambition of obtaining information on the DNA of a single cell[8]. Unlike other techniques that draw their data from thousands of cells, this approach wishes to take into account the heterogeneity of cell populations and the genomic diversity existing between cells in the same organ.
Genomics applied to aging and age-related diseases
We talked about the different techniques of genome analysis, so now what can we do with the data obtained? Sequencing provides thousands of pieces of information that must then be processed. The obvious application is the correlation between genome changes and existing diseases, an approach similar to GWAS. Thanks to sequencing, we can also identify mutations appearing de novo, rare mutations that other techniques cannot discern, or highlight gene variants, giving us new therapeutic perspectives[6].
Genetics, Genomics and Epigenomics
Genetics differs from genomics in its more targeted approach, focusing on a specific gene, its possible mutations and its transmission. Genomics, as we have seen, is concerned with the entire genome and its variations. This type of study has given rise to new areas of research, the most important of which is epigenomics: a mixture of genomics and epigenetics. Like genomics and genetics, epigenomics differs from epigenetics in its subject of study, the genome, or more specifically the epigenome, which represents all changes in the genetic material of a cell (). By combining these approaches, genomic maps are obtained, showing mutations, polymorphisms, genetic variants or DNA methylation rates[9].

Cancer
Several teams around the world have looked at comparing the genome of normal cells to that of cancer cells, particularly in terms of structural variants. A structural variant is a small segment of DNA whose sequence remains the same but which changes conformation (inversion, translocation) giving rise to cellular diversity, beneficial only up to a certain stage[10]. These variants are involved in many diseases, when their rate increases, but remain very difficult to detect. Genomic techniques are the only ones that can identify them, thanks to the use of mathematical algorithms that allow such fine analysis of data that they can find rare nucleotide variants (a few bases and no longer an entire segment)[11, 12]. The development of WGS (whole-genome sequencing) analysis techniques has also made it possible to identify mutations appearing de novo, i.e. not inherited from the parents. Conrad et al. estimated these mutations at about 74 per germ line (the one that gives eggs and sperm)[13]. These mutations are particularly harmful and interesting for sporadic diseases because they are not subject to the natural selection that takes place when a gene passes from one individual to another.
These approaches are very complementary and now make it possible to validate what pathologists saw under the microscope, namely that not all cancer cells are necessarily alike. Several teams have demonstrated this heterogeneity, both phenotypic and genetic[14], by adding a notion of evolution. Indeed, primary tumours and metastases do not seem to have exactly the same genome and adapt to external pressures, such as chemotherapy. This information made it possible to correctly characterize the different types of tumours and predict the progression or relapse of a cancer. Beyond this diagnostic and preventive assistance, genomics also makes it possible to consider ultra-targeted and personalized therapies[15].
Neurodegenerative diseases

Using blood samples, several teams have been working to decipher the genome of neurodegenerative diseases, and more particularly Alzheimer’s disease, the most studied and prevalent of them. Thanks to their work, genes have been identified, notably APOE, CD33 or EPHA1, and their polymorphisms seem to be associated with the development of Alzheimer’s disease. For example, CR1 (chromosome 1) and CLU (chromosome 8) have two loci (a polymorphic zone specific to the gene) very strongly associated with the occurrence of Alzheimer’s disease[17]. In addition to these discoveries, genomics has also made it possible to find previously unknown mutations on “classical” Alzheimer’s disease genes, in particular APP (coding for the amyloid precursor) and MAPT (coding for the Tau protein).
Finally, the most important contribution of genomics to this type of disease is the study of the epigenome, a daughter discipline of genomics. Increasingly, the study of methylation, folding and DNA modification, as well as the study of gene expression regulation phenomena, are crucial tools for understanding the development of neurodegenerative pathologies[18]. For more information, you can visit the NIAGADS website, which lists all public genomic studies in progress. The same type of studies can be found for cardiovascular diseases and metabolic pathologies such as diabetes. All GWAS studies are listed on the site of the National Genome Research Institute and highlight 11 genes for Alzheimer’s disease, 42 genes for cardiovascular disease and 25 genes for diabetes, whose polymorphisms and/or mutations are linked to the occurrence and/or severity of these pathologies.
Towards anti-aging genomics?
Genomics can also be used to determine the propensity of a cell to become senescent and several studies have already shown a difference between senescent phenotypes and others[19]. Aging regulation genes have also been identified, with polymorphisms predisposing to a more or less long lifespan. This is the case of CEBPB, a gene involved in muscle metabolism and whose polymorphisms are risk factors for the occurrence of age-related sarcopenias, which greatly shorten life expectancy[20].
Thanks to all this genomics research, hundreds of researchers have been able to identify common denominators predisposing to age-related diseases. Beyond a diagnostic approach, many of us hope to one day see the emergence of a therapeutic and preventive arsenal in order to manage aging as a whole.
See all our articles on the “-omics” approaches
The “-omics” : a tool to better understand our aging process
What is behind the term “-omics? When we talk of genomics, transcriptomics or proteomics, what are we looking at? Here is a guide to explain it all!
Part 1: Understanding genomics for anti-aging research

How can we not mention genomics and how useful they are. It is the oldest “-omics” approach and still the most studied one even today. It gave birth to whole new concepts, such as epigenetics or systems biology, and opened up the scientific community to new horizons that we had never even dreamed ot!
Part 2: Transcriptomics, a constantly evolving science.

The discovery of non coding RNA led to a Nobel Prize. Enough to say it’s an important field of research! Transcriptomics is allowing the discovery of new tools, new mechanisms and led us to better understand the regulation of transcription.
Part 3: Proteomics, a mish-mash of research fields
 Above all, proteomics are a multidisciplinary approach, taking into account the interactions between fields, namely genomics and transcriptomics. It also refers to different concepts, from immunology to nutrition or cell function.
Above all, proteomics are a multidisciplinary approach, taking into account the interactions between fields, namely genomics and transcriptomics. It also refers to different concepts, from immunology to nutrition or cell function.
Part 4: Metabolomics, the last-born of the “omics”
 Last but not least! Metabolomics is a field that helps us understand very complicated regulation networks and the daily discovery of new cell-cell communication players.
Last but not least! Metabolomics is a field that helps us understand very complicated regulation networks and the daily discovery of new cell-cell communication players.
References
[1] R.W. Holley, et al., Structure of a ribonucleic acid, Science, 1965;147:1462–1465
[2] Sanger, et al., Nucleotide sequence of bacteriophage phi X174 DNA, Nature 1977;265:687–695
[3] F. Sanger, A. Coulson, A rapid method for determining sequences in DNA by primed synthesis with DNA polymerase, J. Mol. Biol. 1975;94:441–448
[4] M. Maxam, W. Gilbert, A new method for sequencing DNA, Proc. Natl. Acad. Sci. U. S. A. 1977;74:560–564
[5] JA. Reuter, DV. Spacek, MP. Snyder, High-Throughput Sequencing Technologies, Molecular Cell, 2015;58(4):Pages 586-597
[6] DC. Koboldt, KM. Steinberg, DE. Larson, RK. Wilson, ER. Mardis, The Next-Generation Sequencing Revolution and Its Impact on Genomics, Cell, 2013;155(1):27-38
[7] Marchini J, Howie B. “Genotype imputation for genome-wide association studies”. Nature Reviews. Genetics. 2010;11(7):499–511
[8] Macaulay IC, Voet T. Single Cell Genomics: Advances and Future Perspectives. Maizels N, ed. PLoS Genetics. 2014;10(1):e1004126
[9] Ben-Avraham D, Muzumdar RH, Atzmon G. Epigenetic genome-wide association methylation in aging and longevity. Epigenomics. 2012;4(5):503-509
[10] Tattini L, D’Aurizio R, Magi A. Detection of Genomic Structural Variants from Next-Generation Sequencing Data. Frontiers in Bioengineering and Biotechnology. 2015;3:92
[11] C.T. Saunders, W.S. Wong, S. Swamy, J. Becq, L.J. Murray, R.K. Cheetham Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs, Bioinformatics, 2012;28:1811-1817
[12] K. Cibulskis, M.S. Lawrence, S.L. Carter, A. Sivachenko, D. Jaffe, C. Sougnez, S. Gabriel, M. Meyerson, E.S. Lander, G. Getz Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples, Nat. Biotechnol., 2013;31:213-219
[13] D.F. Conrad, J.E. Keebler, M.A. DePrist et al., 1000 Genomes Project, Variation in genome-wide mutation rates within and between human families, Nat. Genet., 2011;43:712-714
[14] P.J. Campbell, S. Yachida, L.J. Mudie et al. The patterns and dynamics of genomic instability in metastatic pancreatic cancer, Nature, 2010;467:1109-1113
[15] Uzilov AV, Ding W, Fink MY, et al. Development and clinical application of an integrative genomic approach to personalized cancer therapy. Genome Medicine. 2016;8:627
[16] Antunez C., Boada M., Gonzalez-Perez A., Gayan J., Ramirez-Lorca R., Marin J. The membrane-spanning 4-domains, subfamily A (MS4A) gene cluster contains a common variant associated with Alzheimer’s disease. Genome Med. 2011;3:33
[17] Lambert J.C., Heath S., Even G., Campion D., Sleegers K., Hiltunen M. Genome-wide association study identifies variants at CLU and CR1 associated with Alzheimer’s disease. Nat Genet. 2009;41:1094–1099
[18] Condliffe D., Wong A., Troakes C., Proitsi P., Patel Y., Chouliaras L. Cross-region reduction in 5-hydroxymethylcytosine in Alzheimer’s disease brain. Neurobiol Aging. 2014;35:1850–1854
[19] Pilling LC, Harries LW, Powell J, Llewellyn DJ, Ferrucci L, Melzer D. Genomics and Successful Aging: Grounds for Renewed Optimism? The Journals of Gerontology Series A: Biological Sciences and Medical Sciences. 2012;67A(5):511-519
[20] Hicks GE, Shardell M, Alley DE, Miller RR, Bandinelli S, Guralnik J, Lauretani F, Simonsick EM, Ferrucci L, Absolute strength and loss of strength as predictors of mobility decline in older adults: the InCHIANTI study. J Gerontol A Biol Sci Med Sci. 2012;67(1):66-73
Dr. Marion Tible

Author/Reviewer
Auteure/Relectrice
Marion Tible has a PhD in cellular biology and physiopathology. Formerly a researcher in thematics varying from cardiology to neurodegenerative diseases, she is now part of Long Long Life team and is involved in scientific writing and anti-aging research.
More about the Long Long Life team
Marion Tible est docteur en biologie cellulaire et physiopathologie. Ancienne chercheuse dans des thématiques oscillant de la cardiologie aux maladies neurodégénératives, elle est aujourd’hui impliquée au sein de Long Long Life pour la rédaction scientifique et la recherche contre le vieillissement.
En savoir plus sur l’équipe de Long Long Life
Dr Guilhem Velvé Casquillas

Author/Reviewer
Auteur/Relecteur
Physics PhD, CEO NBIC Valley, CEO Long Long Life, CEO Elvesys Microfluidic Innovation Center
More about the Long Long Life team
Docteur en physique, CEO NBIC Valley, CEO Long Long Life, CEO Elvesys Microfluidic Innovation Center
En savoir plus sur l’équipe de Long Long Life