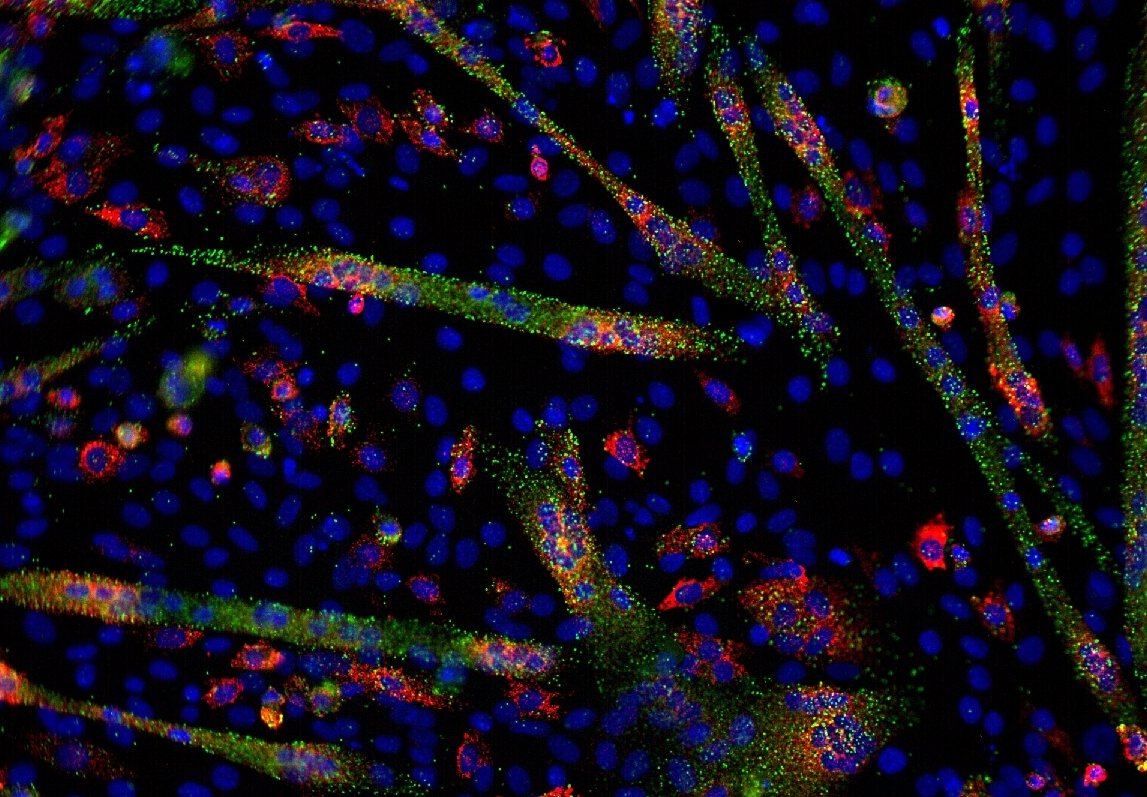
Transcriptomics is the intermediate step between DNA and proteins. However, there are also non-coding RNAs, which do not give proteins, and whose roles are still being studied. All RNAs form what is called the transcriptome. Transcriptomics is a very broad field that will allow the analysis of transcription mechanisms, the regulation of these phenomena and the interactions between RNA, proteins and other factors.
Transcriptomics: How do you study the transcriptome?
Like DNA analysis, RNA analysis combines sequencing techniques. These include sequencing techniques preceded by retrotranscription (to read the DNA sequence), also called RNA-Seq[1] and microarray techniques[2] (like DNA chips but for RNA). These techniques are very similar to those used for DNA (see genomic dossier) and give, beyond sequence information useful for detecting mutations or transcription errors, quantitative information to determine at which level an RNA is expressed. This is called the expression rate of a given RNA: when this rate is high, the DNA will transcribe several copies of RNA, allowing an important expression of proteins. Conversely, when this rate is low, the DNA will be little or not transcribed, resulting in little protein thereafter. These expression rates may vary with time and/or pathologies. Several sites list transcriptome analyses in order to group all transcriptomes according to organs, pathologies, age, treatment or exposure to external factors; you can visit the GEO library or the Expression Atlas for instance.
Quantitative PCR as a transcriptomics tool
Quantitative PCR, or real-time PCR, is a technique dedicated to the study of the transcriptome. Although more tedious than RNAseq approaches or microarrays, it remains the approach of choice for the study of RNA because of its low cost[3].
PCR stands for polymerase chain reaction. This name simply means that the technique is based on the action of a specific enzyme, polymerase, which will allow the synthesis of a complementary strand of DNA.
Given that we’re interested in RNA, it will be necessary to retro-transcribe the RNA into DNA. After this retro-transcription, we obtain a DNA without non-coding regions, corresponding to the gene. Once this DNA has been obtained, PCR makes it possible to monitor the synthesis of its complementary strand using markers (generally fluorochromes). The more RNA there is at the base, the more retrotranscribed DNA there will be and the more quickly the number of complementary strands will increase, thus giving quantitative information on the RNA of interest. In general, 40 so-called “amplification” cycles are performed (i.e. corresponding to a cycle of synthesis of the complementary strands). At a given moment, the signal emitted by the complementary strands will pass above a certain limit, giving a value called Ct (for “cycle threshold”).
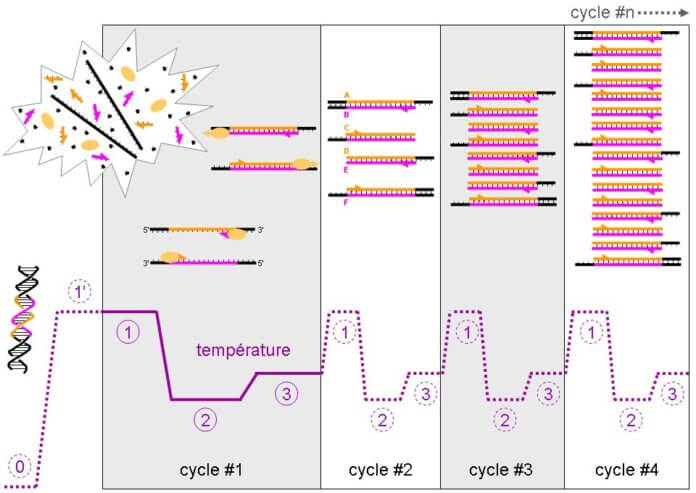
The smaller this value, the faster the synthesis of the complementary strands will be, meaning that the number of RNA of interest was originally high and that the protein it will give will be strongly expressed[3].
This technique is also applicable for non-coding RNAs but will obviously not give information on the future expression rate of a protein, because a non-coding RNA does not code for any protein, as its name indicates. Moreover, the notion of expression of a protein from transcriptomic data is complex because translation (RNA to protein) can also be regulated up or down. The most important concept is therefore purely quantitative, with a RNA expression rate reflecting gene activity and possibly protein synthesis.
What do we do with transcriptomics data?
Measuring gene expression in different tissues, ages or pathophysiological conditions, as is done through the study of the transcriptome, provides a better understanding of how a given gene will be regulated and provides important information about the function and needs of a given organ[4]. Transcriptomics can also be used to find new functions for a gene and/or discover new genes. All these applications obviously lead to a better understanding of diseases, how they work and their genetic risk factors[4]. In parallel to this information, transcriptomics also allows an analysis of all RNAs, including the coding RNAs we have mentioned (or messenger RNA or mRNA), but also non-coding RNAs.
Non-coding NRAs: prospects for the future of transcriptomics
It is now established that about 70% of the genome is transcribed into RNA but only 2% will give proteins[5]. The non-coding RNAs are thus largely in the majority and include[6]:
- miRNAs, or microRNAs, small RNAs of about 20 base pairs that have a major role in gene expression regulation. Indeed, they can bind to the mRNA, preventing its translation into protein by different mechanisms: either the mRNA is destroyed, or the mRNA is destabilized, or the mRNA still goes to the ribosome but will not be able to translate the protein correctly.
- tRNAs, known as transfer RNAs, whose role is to link amino acids to mRNAs within ribosomes during translation. Therefore, tRNAs are necessary for the translation and biosynthesis of proteins in accordance with the genetic sequence.
- ribosomal RNAs, contained in ribosomes and whose role is central in translation. These rRNAs are among the oldest and are extremely well preserved between species. They are very often used as a reference during transcriptomic analyses.
- siRNAs, or small-interfering RNAs, similar to miRNAs but operating in different signaling pathways. They have an important role in the regulation of translation because their association with mRNA leads to their destruction (and the absence of protein production). Their synthetic form is widely used in the pharmaceutical industry and research.
- piwiRNAs or piRNAs (for “piwi-interfering”), which interact with piwi proteins, regulatory proteins capable of binding to RNA. When piwiRNs form a complex with their target proteins, they can regulate epigenetic and post-transcriptional phenomena.
- snoRNAs (for Small-NucleOlar), small RNAs associated with modifications (methylation, among others) of other RNAs.
- snRNAs (for “small nuclear”), which help in the synthesis of mRNAs.
- exRNAs (for “exosomal”), small RNAs or portions of RNA present outside cells in what are called exosomes (small vesicles).
- lncRNAs (for “long non coding”), longer non-coding RNAs (about 200 base pairs), whose function is still being discussed.
The discovery of non-coding RNAs, particularly interfering RNAs, earned Professor Fire and Professor Mello the Nobel Prize in Physiology and Medicine in 2006. The reason for this award is simple: their discovery was crucial to understanding mRNA regulation, a relatively unclear process. MiRNAs and siRNAs are currently the most studied because they have the ability to send a degradation signal to the cell to destroy the mRNAs they regulate. Moreover, they have been found in blood, making them important biomarkers of biological processes and pathologies[7].
These circulating miRNAs should be good biomarkers of certain diseases and physiological conditions. They are also new avenues for developing drugs capable of regulating protein synthesis and perhaps helping to cure these age-related pathologies. Because of their size and ability to cross the blood-brain barrier, they are also good therapeutic candidates themselves. In cancer, miRNAs and lncRNAs are extremely promising, with significant deregulations[9, 10]. The injection of miR-16 in mouse models of prostate cancer demonstrated, in particular, the downward regulation of the miRNA target and a decrease in tumour mass[11].
These circulating miRNAs should be good biomarkers of certain diseases and physiological conditions. They are also new avenues for developing drugs capable of regulating protein synthesis and perhaps helping to cure these age-related pathologies. Because of their size and ability to cross the blood-brain barrier, they are also good therapeutic candidates themselves. In cancer, miRNAs and lncRNAs are extremely promising, with significant deregulations[9, 10]. The injection of miR-16 in mouse models of prostate cancer demonstrated, in particular, the downward regulation of the miRNA target and a decrease in tumour mass[11].
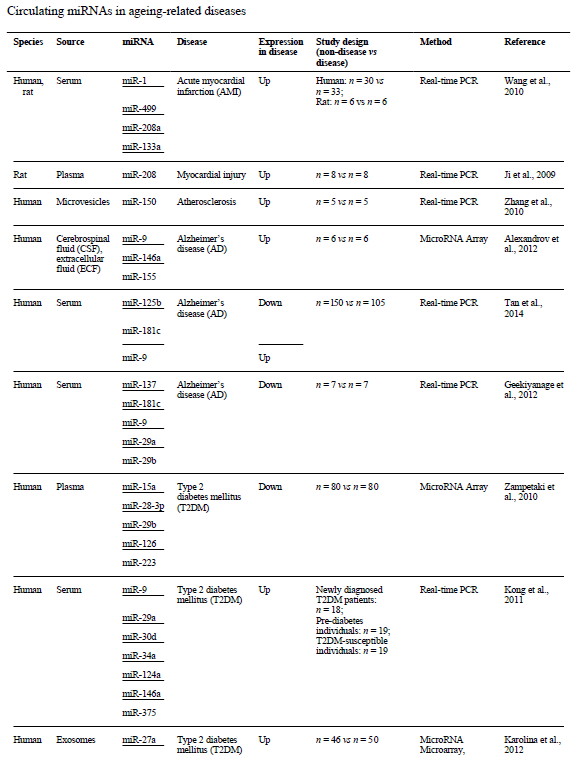
Transcriptomics in the fight against aging
Beyond the role of non-coding RNAs as biomarkers and therapeutic targets, they are also RNAs that can be regulated over a lifespan. For example, telomerase, an enzyme that adds protective segments to our chromosomes to prevent the loss of information during cell divisions (see our telomeres articles), is called reverse transcriptase. This type of enzyme is capable of retrotranscribing RNA into DNA and this is what happens at the telomeric level: telomerase carries specific RNA that it will add at the end of our chromosomes once transformed into DNA[12]. These non-coding RNAs are central protectors of our genome and their level decreases with age, as does telomerase activity.
In parallel, the global transcriptomic profiles (not only non-coding RNAs) of our cells also vary during aging. In our immune cells, RNAs involved in the inhibition of autophagy (cell detoxification system) were increased while those involved in the regulation of oxidative phosphorylation (oxidant production), cytoskeletal synthesis or insulin response were decreased[13]. In the brain, and more particularly the grey matter, the mRNAs of the proliferation, adhesion and cellular differentiation pathways were regulated down in parallel with an increase in mRNAs involved in the inflammatory pathways, suggesting a decrease in cellular regeneration processes and an increase in neuro-inflammation with age[14, 15]. It also appears that these changes occur more rapidly in women, probably explaining the higher prevalence of neurodegenerative diseases in this segment of the population[16].
New discoveries are being made every day and more and more teams are interested in aging as such. A study in 2014 has notably launched a study of the blood transcriptome of more than 2500 people in the context of physiological aging (without associated pathologies). They found that the pathways of T-cell activation, RNA elongation during transcription, cytolysis and DNA-related metabolic processes were all altered and that these changes were associated with life expectancy[17]. In mice, another research team looked at the difference between physiological aging and pathological aging. Biopsies of five organs – liver, kidney, spleen, lung and brain – were collected regularly throughout the life of the mice and their transcriptomes were studied. Over time, researchers have highlighted changes linked to aging, sometimes accentuated or supplemented by changes linked to pathologies that appear with age[18].
All these combined approaches offer a complementary view of genomic data and make it possible to understand aging from different angles: both to implement diagnostic and therapeutic tools, but also to discover new pathways or new genes involved in changes in our body and its functions over the course of our lives.
See all our articles on the “-omics” approaches
The “-omics” : a tool to better understand our aging process
What is behind the term “-omics? When we talk of genomics, transcriptomics or proteomics, what are we looking at? Here is a guide to explain it all!
Part 1: Understanding genomics for anti-aging research

How can we not mention genomics and how useful they are. It is the oldest “-omics” approach and still the most studied one even today. It gave birth to whole new concepts, such as epigenetics or systems biology, and opened up the scientific community to new horizons that we had never even dreamed ot!
Part 2: Transcriptomics, a constantly evolving science.

The discovery of non coding RNA led to a Nobel Prize. Enough to say it’s an important field of research! Transcriptomics is allowing the discovery of new tools, new mechanisms and led us to better understand the regulation of transcription.
Part 3: Proteomics, a mish-mash of research fields
 Above all, proteomics are a multidisciplinary approach, taking into account the interactions between fields, namely genomics and transcriptomics. It also refers to different concepts, from immunology to nutrition or cell function.
Above all, proteomics are a multidisciplinary approach, taking into account the interactions between fields, namely genomics and transcriptomics. It also refers to different concepts, from immunology to nutrition or cell function.
Part 4: Metabolomics, the last-born of the “omics”
 Last but not least! Metabolomics is a field that helps us understand very complicated regulation networks and the daily discovery of new cell-cell communication players.
Last but not least! Metabolomics is a field that helps us understand very complicated regulation networks and the daily discovery of new cell-cell communication players.
References :
[1] Wang Z, Gerstein M, Snyder M, RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet. 2009; 10(1):57-63
[2] Schena M, Shalon D, Davis RW, Brown PO, Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science. 1995; 270(5235):467-70
[3] Alwine JC, Kemp DJ, Stark GR, Method for detection of specific RNAs in agarose gels by transfer to diazobenzyloxymethyl-paper and hybridization with DNA probes. Proc Natl Acad Sci U S A. 1977; 74(12):5350-4
[4] Han Y, Gao S, Muegge K, Zhang W, Zhou B. Advanced Applications of RNA Sequencing and Challenges. Bioinformatics and Biology Insights. 2015;9(Suppl 1):29-46
[5] Bertone P, Stolc V, Royce TE, Rozowsky JS, Urban AE, Zhu X, Rinn JL, Tongprasit W, Samanta M, Weissman S, Gerstein M, Snyder M, Global identification of human transcribed sequences with genome tiling arrays. Science. 2004; 306(5705):2242-6
[6] Hirose T, Mishima Y, Tomari Y. Elements and machinery of non-coding RNAs: toward their taxonomy. EMBO Reports. 2014;15(5):489-507
[7] Ling H, Fabbri M, Calin GA. MicroRNAs and other non-coding RNAs as targets for anticancer drug development. Nature reviews Drug discovery. 2013;12(11):847-865
[8] Jung HJ, Suh Y. Circulating miRNAs in Ageing and Ageing-Related Diseases. Journal of genetics and genomics. 2014;41(9):465-472
[9] Gutschner T, Diederichs S. The hallmarks of cancer: A long non-coding RNA point of view. RNA Biology. 2012;9(6):703-719
[10] Berindan-Neagoe I, Monroig P, Pasculli B, Calin GA. MicroRNAome genome: a treasure for cancer diagnosis and therapy. CA: a cancer journal for clinicians. 2014;64(5):311-336
[11] Iguchi H, Kosaka N, Ochiya T, Secretory microRNAs as a versatile communication tool. Commun Integr Biol. 2010; 3(5):478-81
[12] Ly H, Blackburn EH, Parslow TG. Comprehensive Structure-Function Analysis of the Core Domain of Human Telomerase RNA. Molecular and Cellular Biology. 2003;23(19):6849-6856
[13] Reynolds LM, Ding J, Taylor JR, et al. Transcriptomic profiles of aging in purified human immune cells. BMC Genomics. 2015;16(1):333
[14] Kochunov P, Charlesworth J, Winkler A, et al. Transcriptomics of cortical gray matter thickness decline during normal aging. NeuroImage. 2013;0:273-283
[15] Kang HJ, Kawasawa YI, Cheng F, et al. Spatiotemporal transcriptome of the human brain. Nature. 2011;478(7370):483-489
[16] Yuan Y, Chen Y-PP, Boyd-Kirkup J, Khaitovich P, Somel M. Accelerated aging-related transcriptome changes in the female prefrontal cortex. Aging Cell. 2012;11(5):894-901
[17] Van den Akker EB, Passtoors WM, Jansen R, et al. Meta-analysis on blood transcriptomic studies identifies consistently coexpressed protein–protein interaction modules as robust markers of human aging. Aging Cell. 2014;13(2):216-225
[18] Melis JPM, Jonker MJ, Vijg J, Hoeijmakers JHJ, Breit TM, van Steeg H. Aging on a different scale – chronological versus pathology-related aging. Aging (Albany NY). 2013;5(10):782-788
Dr. Marion Tible

Author/Reviewer
Auteure/Relectrice
Marion Tible has a PhD in cellular biology and physiopathology. Formerly a researcher in thematics varying from cardiology to neurodegenerative diseases, she is now part of Long Long Life team and is involved in scientific writing and anti-aging research.
More about the Long Long Life team
Marion Tible est docteur en biologie cellulaire et physiopathologie. Ancienne chercheuse dans des thématiques oscillant de la cardiologie aux maladies neurodégénératives, elle est aujourd’hui impliquée au sein de Long Long Life pour la rédaction scientifique et la recherche contre le vieillissement.
En savoir plus sur l’équipe de Long Long Life
Dr Guilhem Velvé Casquillas

Author/Reviewer
Auteur/Relecteur
Physics PhD, CEO NBIC Valley, CEO Long Long Life, CEO Elvesys Microfluidic Innovation Center
More about the Long Long Life team
Docteur en physique, CEO NBIC Valley, CEO Long Long Life, CEO Elvesys Microfluidic Innovation Center
En savoir plus sur l’équipe de Long Long Life