DNA methylation: a powerful epigenetic regulator
Throughout our lives, our DNA undergoes modifications linked to our diet, environment, diseases and adaptative phenomenons. They are called epigenetic modifications, and they are closely related to aging. Breakthroughs in quantitative methods for full genome analysis (genome-wide associated studies, or GWAS) allowed to shed some light on epigenetic modification and to study how they are regulated.
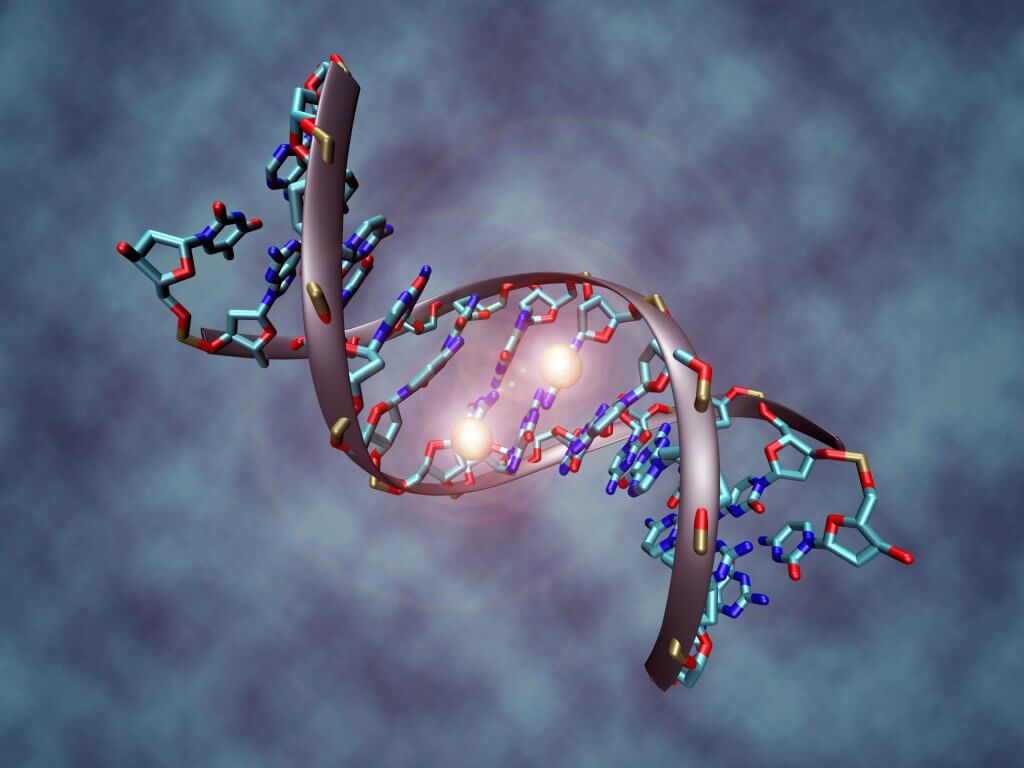
Epigenetic modifications are diverse and can happen at any time during a lifetime, as a reply to signals that our cells receive: stress, smoking, diet, hormonal response… They can be temporary and help the body face a specific emergency, or long-lasting, in which case they have a direct impact on gene expression and long-term DNA integrity. They can take many forms and result from the work of specific enzymes that add small structures to our DNA, either directly or through DNA-associated proteins like histones. The modifications we know best today are also the most common : glycosylation (sugar addition), methylation (addition of a methyl group on our DNA), and histone methylation and acetylation.
They have a direct impact on gene transcription (and full-protein generation) because they can block important DNA sites, as well as the access to sequences that are indispensable to transcription. Methylation can lead to the inactivation of a given gene. The alterations caused by aging-related methylation are called “epigenetic derivations”, a number of events that lead to the gradual de-methylation of the genome, and the hyper-methylation or certain areas called CpG islands [1].
Epigenetics and diseases
Epigenetic anomalies can help diseases develop. This was proven for cancers [2], and more recently in neurodegenerative diseases [3, 4, 5] and metabolic diseases such as diabetes [6].
The methylation process regulates many others, such as cell differentiation, division, multiplication and survival. These phenomenons allow to regulate the expression of some genes, and when a malfunction occurs, it can activate genes that were not supposed to be activated (such as oncogenes) or inhibit other genes that were supposed to stay active (such as tumor suppressors).
Aging and methylation
Methylation is a specific type of epigenetic modification: it happens at the cytosine level of our DNA, one of the four DNA bases, with adenine, guanine and thymine. One family of enzymes, the DNA methyltransferase (DNMTs) allow the addition of a methyl group (of chemical formula CH3) on one of the carbones that make up the cytosine structure [7,8]. This methylation play a major role in embryongenesis, and later in life, in genetic regulation. It is responsible for the selective expression of paternal and maternal genes, a mechanism called “genomic imprinting” which allows to inactivate one of the two X chromosomes in female embryos. After this developmental phase, methylation will happen in about 1% of the cytosines that make up the DNA, most frequently inside the CpG islands [9].
During aging, the DNMTs start the methylation process more and more often, in some cytosine-rich areas that are paramount to good gene transcription: inside the promoters’ CpG islands. These are non-coding areas (they contain no genetic information) located ahead of the genes and which will welcome the RNA polymerase, an enzyme family in charge of transcription. When the promoters are hyper-methylated, the RNA polymerase cannot latch on and this inhibits gene transcription [1, 8, 10].
This is why there is a link between genome regulation, methylation and aging. Many studies have shown that the levels of 5-methyl-cytosine (5mC) was correlated with age [11, 12] which gives us additional tools in the metrology of aging.
Measuring methylation: clues about your biological age?
There are tools, similar to those used to decipher our genome, to get comprehensive information about the epigenetic modifications of our DNA; they are about our epigenome.
Methylation can be measured with those techniques, such as Illumina (Infinium HumanMethylation BeadChip assays), a process to obtain reliable information over the number of CpG islands and how many were methylated, over a whole genome.
A research team managed to identify 393 sequences in the CpG islands that could predict the biological age of patients, over a population of more than 2,000 people and 8,000 samples from different tissues. The double interest of these markers is that they are highly sensitive (96% reliable) and very specific to the tissue they are tested in [13]. More recently, a study limited to blood predicted the biological age of a person from only three specific CpG sites, showing the “epigenetic clock” theory to be true [14].
To the future: “epimedicine” against methylation
Assuming that epigenetic modifications are usually reversible, many hopes are placed in targeted therapies to correct these changes. The idea led us to the development of epimedicine, which can eliminate abnormal DNA modifications. Two families of molecules are already being developed: DNA methyltransferase inhibitors (DNMTi) and histone deacetylase inhibitors (HDACi).
Sadly, for now, their action is not specific enough for a targeted effect without side effects. However, many research teams are currently working on the amelioration of current molecules, and even multi-therapies that associate epimedicine and regular molecules.
All our articles on the Metrology of Aging
Measuring aging: what is your biological age?

Measuring our biological age and take consistent dispositions to avoid it, it’s a dream come true !
Part 1: Measuring aging: Cartographie PhysiologiqueⓇ
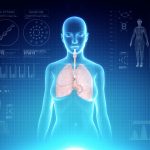 This method to measure aging and physiological age measures the many physical and biological characteristics that were definitely linked to the aging process (cholesterol levels, unipedal stance test, central obesity…)
This method to measure aging and physiological age measures the many physical and biological characteristics that were definitely linked to the aging process (cholesterol levels, unipedal stance test, central obesity…)
Part 2: Measuring aging: the eighteen markers of Duke University

This method follows the decline of many organic systems (cardiovascular, lungs, kidneys, liver, teeth, and immune system) by quantifying 18 markers for chronic diseases linked to aging. This technique was developed by a research team from Duke University, under the supervision of Dr. Daniel Belsky.
Part 3: DNA methylation, a promising lead towards the measure of aging?
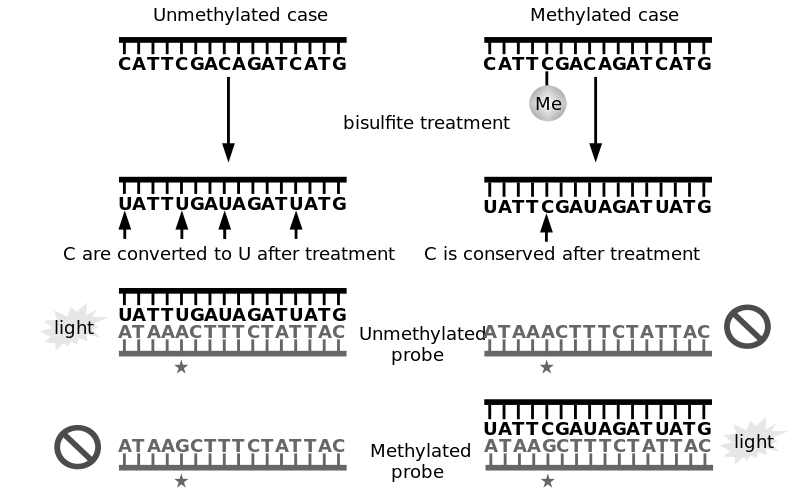
 DNA methylation levels grow with the number of divisions a cell undergoes. The number of cell divisions grows with time. Therefore, measuring the methylation of DNA helps to quantify aging. Treating DNA with bisulfites allows to convert non-methylated cytosines into uraciles, so the methylated cytosines can easily be identified. We deduce a physiological age from their numbers.
DNA methylation levels grow with the number of divisions a cell undergoes. The number of cell divisions grows with time. Therefore, measuring the methylation of DNA helps to quantify aging. Treating DNA with bisulfites allows to convert non-methylated cytosines into uraciles, so the methylated cytosines can easily be identified. We deduce a physiological age from their numbers.
Part 4: Telomeres, at witnesses of the physiological age, as a tool to measure aging
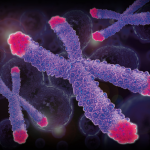 Telomeres can be found at the end of our chromosomes, and get shorter with age. There are many techniques to measure their length (TRFs, Q-Fish, qPCR, STELA…) based on the Southern Blot principle, PCR and in situ hybridization.
Telomeres can be found at the end of our chromosomes, and get shorter with age. There are many techniques to measure their length (TRFs, Q-Fish, qPCR, STELA…) based on the Southern Blot principle, PCR and in situ hybridization.
References
[1] Jones MJ, Goodman SJ, Kobor MS. DNA methylation and healthy human aging. Aging Cell. 2015;14(6):924-932
[2] Weisenberger DJ. Characterizing DNA methylation alterations from The Cancer Genome Atlas. The Journal of Clinical Investigation. 2014;124(1):17-23
[3] Wüllner U, Kaut O, deBoni L, Piston D, Schmitt I. DNA methylation in Parkinson’s disease. J Neurochem. 2016 Oct;139 Suppl 1:108-120
[4] Horvath S, Zhang Y, Langfelder P, et al. Aging effects on DNA methylation modules in human brain and blood tissue. Genome Biology. 2012;13(10):R97
[5] Xu X. DNA methylation and cognitive aging. Oncotarget. 2015;6(16):13922-13932.
[6] Chambers JC, Loh M, Lehne B, et al. Epigenome-wide association of DNA methylation markers in peripheral blood from Indian Asians and Europeans with incident type 2 diabetes: a nested case-control study. The lancet Diabetes & endocrinology. 2015;3(7):526-534
[7] Johnson AA, Akman K, Calimport SRG, Wuttke D, Stolzing A, de Magalhães JP. The Role of DNA Methylation in Aging, Rejuvenation, and Age-Related Disease. Rejuvenation Research. 2012;15(5):483-494
[8] Zampieri M., Ciccarone F., Calabrese R. , Franceschi C., Burkle A., Caiafa P., Reconfiguration of DNA methylation in aging. Mech. of Ageing and Dev., 2015, vol 151; P. 60-70
[9] Ehrlich M, Gama-Sosa MA, Huang L-H, Midgett RM, Kuo KC, McCune RA, Gehrke C. Amount and distribution of 5-methylcytosine in human DNA from different types of tissues or cells. Nucleic Acids Res. 1982;10:2709–21.
[10] Jung M, Pfeifer GP. Aging and DNA methylation. BMC Biology. 2015;13:7
[11] Wilson VL, Smith RA, Ma S, Cutler RG. Genomic 5-methyldeoxycytidine decreases with age. J Biol Chem. 1987;262:9948–51
[12] Wilson VL, Jones PA. DNA methylation decreases in aging but not in immortal cells. Science. 1983;220:1055–7
[13] Horvath S. DNA methylation age of human tissues and cell types. Genome Biol. 2013;14:R115
[14] Weidner CI, Lin Q, Koch CM, Eisele L, Beier F, Ziegler P, et al. Aging of blood can be tracked by DNA methylation changes at just three CpG sites. Genome Biol. 2014;15:R24
Farah Bahou

Author
Auteure
Farah studied biochemistry, therapeutics and molecular and biopharmaceutical innovation at Aix-Marseille university and Paris 7 Diderot university.
More about the Long Long Life team
Farah a étudié la biochimie, la thérapeutique et les innovations moléculaires et biopharmaceutiques à l’université d’Aix-Marseille et à l’université Paris 7 Diderot.
En savoir plus sur l’équipe de Long Long Life
Dr Guilhem Velvé Casquillas

Author/Reviewer
Auteur/Relecteur
Physics PhD, CEO NBIC Valley, CEO Long Long Life, CEO Elvesys Microfluidic Innovation Center
More about the Long Long Life team
Docteur en physique, CEO NBIC Valley, CEO Long Long Life, CEO Elvesys Microfluidic Innovation Center
En savoir plus sur l’équipe de Long Long Life


![[Video] Eurosymposium on Healthy Ageing, Brussels, 2018 Eurosymposium on Healthy Aging](https://www.longlonglife.org/wp-content/uploads/2019/07/P1310252-218x150.jpg)