Epigenetic alterations as causes of aging
Aging can be defined as all the phenomena that mark the evolution of a living organism towards death[1]. These lead to a progressive loss of physiological integrity, which eventually alters biological functions[2]. Over time, aging is expressed by the occurrence of typical diseases such as cancer, type 2 diabetes, cardiovascular disorders and neurodegenerative diseases. Today, it is recognized that the biological phenomena responsible for aging include at least nine molecular and cellular processes (see article: Understanding the biological causes of aging and longevity). These causes of aging are the subject of numerous scientific publications, including the excellent article “The Hallmarks of Aging”[2], and the number of research projects on these mechanisms continues to increase.
All these biological phenomena deserve greater attention to understand their complexity. As we did previously by devoting a series of articles to telomere shortening, this article will deal specifically with epigenetic alterations as causes of aging. This is a complex process because it involves many biological mechanisms that, in combination, will affect the normal pattern of epigenetic processes.
Understanding genomics to understand epigenetic alterations
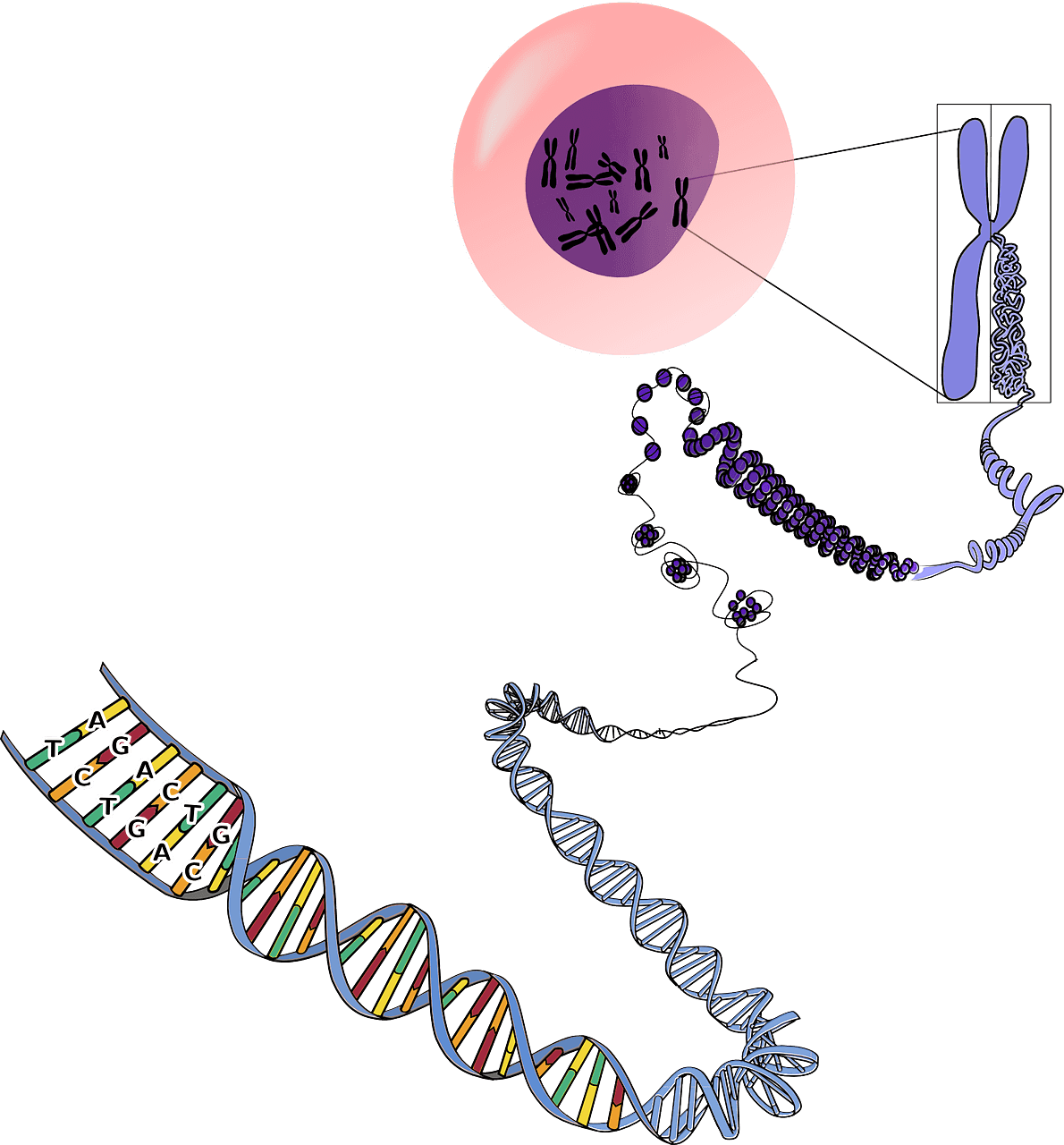
To understand the concept of epigenetics, we must first understand what it affects: the genome. Our genetic information is carried by DNA. It is a four-letter code, A, T, G and C, also called nucleotides, or bases, and the chain of which forms a sequence (a gene is a part of a sequence). DNA interacts with a multitude of biological elements, among others to transcribe it. During transcription, the DNA sequence is copied to form RNA. Then the RNA is read and “translated” into proteins, this is the translation stage. So when a gene is transcribed and then translated, it is said to be expressed.
The human genome contains more than 3 billion base pairs. Such a quantity of nucleotides placed end to end results in a sequence length far too large to enter the cell nucleus. To counter this problem, DNA is associated with proteins, including histones around which DNA wraps to form what is called a nucleosome. Thus, DNA is compacted as chromatin and can be contained in the cell nucleus. There are two forms of chromatin: a highly condensed form called heterochromatin and a less condensed form called euchromatin. The latter is accessible to all transcriptional machinery, so it is in this form that gene expression can take place.
What about epigenetic alterations?
Epigenetics is the study of the mechanisms that modify the state of chromatin, thus allowing the expression of genes to be modulated without changing the sequence. These mechanisms include DNA methylation, histone modifications, chromatin remodelling and transcriptional alterations of specific RNAs.
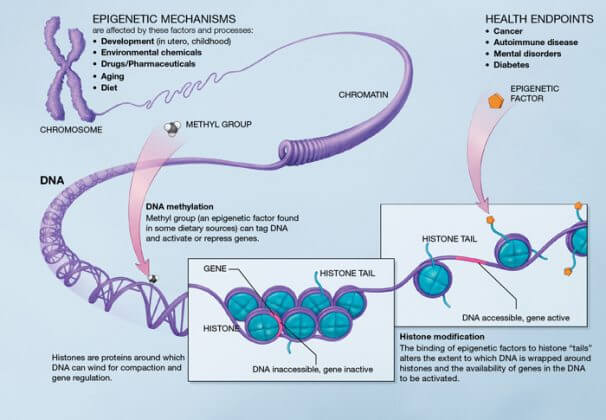
DNA methylation
Methylation levels vary throughout life, but with age they tend to decrease overall. Methylation is a phenomenon in which a methyl chemical group (-CH3) is added to a cytosine base (C) of DNA. It causes DNA condensation, the form in which genes are not transcribed.
Gene modulation is an essential process for maintaining cellular balance. Indeed, all the cells of the body carry the same genome, but they do not all have the same purpose. As a result, a liver cell does not synthesize the same proteins as a skin cell. The expression of certain genes must therefore be activated or repressed, depending on the fate of the cell. Age-related methylation changes occur either randomly on certain cytosines (“Epigenetic drift”) or on specific regions of DNA where methylation changes are associated with age. The latter are used to measure aging.
DNA methylation at the root of the epigenetic clock principle
DNA methylation has been used in recent years as a reliable measurement tool for estimating biological age. This phenomenon, called the epigenetic clock, is based on CpG sites (DNA portion of two bases, the first of which is a cytosine and the second a guanine and linked together by a phosphate bond) associated with age and whose methylation profile can be used as an accurate indicator of biological age[6].

The difference between the age predicted by the epigenetic clock (our biological age) and chronological age is called accelerated aging and may be associated with an increased risk of disease or mortality. In other words, if the biological age is higher than the chronological age, the risk of developing age-related diseases will be higher. Accelerated aging has been shown to be associated with a high risk of cancer and decreased survival during and after chemotherapy[7], as well as increased risk of mortality from cardiovascular disease[8].
Epigenetic alterations through histone modifications
As we age, histones, like DNA, undergo epigenetic changes. Histones are nucleosomes, protein complexes around which DNA wraps to condense or decondense chromatin. The modifications that affect them are mainly methylations and acetylations (addition of an acetyl chemical group -COCH3). Methylation aberrations would lead to an increase in cancer recurrence and a low survival rate. With age, changes in methylation are accompanied by an overall loss in the acetylation level of some histones (a process called hypoacetylation). Studies in mice have shown that preventing age-related hypoacetylation prevents cognitive impairment and reduces the severity of conditions such as Parkinson’s disease, osteoporosis and stroke[9,10].
Epigenetic alterations by chromatin remodeling
To maintain chromatin architecture, chromosomal proteins such as HP1α and chromatin remodeling factors such as Polycomb proteins and the NuRD complex are essential. With age, the level of these proteins decreases and induces an overall loss of heterochromatin, established as an aging mark[2]. This phenomenon is found in people with premature aging syndromes, such as Werner’s disease and Huntchinson-Gilford disease[11].
Epigenetic alterations by transcriptional modification
Transcriptional alterations are all changes in non-coding RNA levels. These are promising tools for the development of new diagnostic or prognostic biomarkers, but also for the development of new therapeutic targets. As their name suggests, they are RNAs that are not translated into proteins, but they are important regulators of transcription and chromatin architecture. These RNAs include short non-coding RNAs and long non-coding RNAs (lncRNAs). These RNAs are transcribed aberrantly in cancers and cardiovascular diseases, among others.
Reversibility of epigenetic alterations
Unlike DNA mutations, epigenetic alterations are reversible. Indeed, epigenetic changes are introduced by enzymes and their actions are modulable. In addition, the signalling pathways regulating these enzymes can be targeted by drugs. Finally, changes in behaviour and lifestyle can also modify epigenetic alterations.
Epigenome editing
This technique alters the epigenetic states of chromatin as follows: factors bind to specific DNA sites to induce or suppress epigenetic alterations. Recently, epigenome editing technology based on CRISPR-Cas9 has been used to target different genomic sites to reverse epigenetic alterations[12]. Epigenome editing technology increases research possibilities and could lead to the development of new therapeutic targets and treatments. However, it remains uncertain on some points: the results obtained may be limited because other causes come into play in the development of diseases.
Definitions :
Epigenetics: the study of chromatin modification mechanisms to modulate gene expression without changing the nucleotide sequence of DNA.
Histone: the constituent proteins of the nucleosome, a protein complex around which the chromatin is wound to condense or decondense.
CpG dinucleotide: a CpG site is a DNA portion of two bases, the first of which is a cytosine and the second a guanine and which are linked together by a phosphate bond.
Homeostasis: This is a state in which functions are properly performed leading to the proper organization of a system.
Chronological age: this is the age given by the date of birth, not to be confused with the biological age given to your cells and based on the functioning of different cellular activities.
Chromosomal protein: protein that interacts with DNA and regulates chromatin condensation. Their action condenses chromatin and differs from that of histones which is also mechanical.
Chromatin remodeling factor: these are protein complexes that modify the structure of the nucleosome to modify the state of condensation of the chromatin.
Biomarker: This is a measurable characteristic that can detect a normal or pathological biological process.
References:
[1] http://www.larousse.fr/dictionnaires/francais/vieillissement/81927
[2] Carlos Lopez-Otin, Maria A. Blasco, Linda Partridge, Manuel Serrano and Guido Kroemer. The Hallmarks of Aging, Cell 153, June 2013, 1194-1217.
[3] Neelam Goel, Priya Karir, Vivek Kumar Garg. Role of DNA methylation in human age prediction, Mechanisms of Ageing and Development 166 (2017) 33-41.
[4] Bérénice A. Benayoun, Elizabeth A. Pollina and Anne Brunet. Epigenetic regulation of ageing: linking environmental inputs to genomic stability, Nature Reviews, Molecular Cell Biology, volume 16 (October 2015), 593-610.
[5] Ciccarone, F., Mechanisms of Ageing and Development (2017), https://doi.org/10.1016/j.mad.2017.12.002
[6] Declerck, K., Mechanisms of Ageing and Development (2018), https://doi.org/10.1016/j.mad.2018.01.002
[7] Pierre-Antoine Dugué, Julie K. Bassett, JiHoon E. Joo, Chol-Hee Jung, Ee Ming Wong, Margarita Moreno-Betancur, Daniel Schmidt, Enes Makalic, Shuai Li, Gianluca Severi, Allison M. Hodge, Daniel D. Buchanan, Dallas R. English, John L. Hopper, Melissa C. Southey, Graham G. Giles and Roger L. Milne. DNA methylation-based biological aging and cancer risk and survival: Pooled analysis of seven prospective studies. Int. J. Cancer: 00, 00–00 (2017).
[8] Laura Perna, Yan Zhang, Ute Mons, Bernd Holleczek, Kai-Uwe Saum and Hermann Brenner. Epigenetic age acceleration predicts cancer, cardiovascular, and all-cause mortality in a German case cohort, Clinical Epigenetics (2016) 8:64.
[9] Nard Kubben and Tom Misteli. Shared molecular and cellular mechanisms of premature ageing and ageing-associated diseases, Nature Reviews, Molecular Cell Biology, Volume 18 (October 2017), 595-609.
[10] Ibraheem Ali, Ryan J. Conrad, Eric Verdin and Melanie Ott. Lysine Acetylation Goes Global: From Epigenetics to Metabolism and Therapeutics, Chem. Rev. 2018, 118, 1216−1252.
[11] Lauren N. Booth and Anne Brunet. The Aging Epigenome, Mol Cell. 2016 June 2; 62(5): 728–744. doi:10.1016/j.molcel.2016.05.013.
[12] Cia-Hin Lau and Yousin Suh. Genome and Epigenome Editing in Mechanistic Studies of Human Aging and Aging-Related Disease, Gerontology. 2017 ; 63(2): 103–117. doi:10.1159/000452972.
Anne Fischer

Author
Auteur
Anne is studying medicine science at the Institute of Pharmaceutical and Biological Science in Lyon and she has graduated with a Bachelor’s degree in molecular and cellular biology at the University of Strasbourg.
More about the Long Long Life team
Anne étudie les sciences du médicament à l’Institut des Sciences Pharmaceutiques et Biologiques de Lyon. Elle est titulaire d’une licence en biologie moléculaire et cellulaire de l’Université de Strasbourg.
En savoir plus sur l’équipe de Long Long Life
Dr Guilhem Velvé Casquillas

Author/Reviewer
Auteur/Relecteur
Physics PhD, CEO NBIC Valley, CEO Long Long Life, CEO Elvesys Microfluidic Innovation Center
More about the Long Long Life team
Docteur en physique, CEO NBIC Valley, CEO Long Long Life, CEO Elvesys Microfluidic Innovation Center
En savoir plus sur l’équipe de Long Long Life